Bifidobacterium deficit in United States infants drives prevalent gut dysbiosis
Study population
The IRB-approved My Baby Biome study (Western Consulting Group IRB protocol PBI-2022-01; NCT05472688) was conducted as a decentralized study. Individuals were recruited to the study throughout the United States primarily through social media. After a brief questionnaire to determine eligibility, caregivers of infants roughly between the ages of one month and two months signed an informed consent form and were subsequently sent a fecal collection kit. The study cohort was selected such that it had racial and ethnic diversity comparable to the United States as well as representative levels of birth and feeding modes. Caregivers were instructed to return the fecal collection kit before the child turned 3 months old and completed an in-depth questionnaire at the time of stool collection. Additional questionnaires were collected at six months, one year, and two years and will continue to be collected on a yearly basis up until the 7th birthday.
Stool collection
For the initial collection, fecal samples were collected at home with stool sampling kits that contained Covidien Precision Stool Collector cups and spoons, ice packs to keep the sample cool, and a DNA/RNA Shield Fecal Collection Tube (Zymo Research). Initial samples were scooped immediately from the infant’s diaper and were express shipped overnight to the lab. Scooped samples were both raw fecal samples and DNA/RNA Shield Fecal Collection Tube samples. DNA/RNA Shield Fecal Collection Tubes were stored at −20 °C upon arrival. Raw fecal samples were processed upon arrival in an anaerobic chamber if they were at temperatures less than 17 °C. Prior to processing, color, weight, and consistency of the stool was noted. Phosphate-buffered saline (PBS) aliquots were generated for analysis. A separate aliquot was taken to measure the pH of the sample. Aliquots were labeled and frozen at −80 °C for long-term storage. PBS aliquots were used for whole-genome sequencing, or if the sample was over the appropriate temperature, the DNA/RNA tube was processed instead. Statistical analysis revealed that the method of DNA collection and isolation (raw fecal versus DNA/RNA tubes) had a significant impact on microbial composition (PERMANOVA, p = 0.02). Consequently, we adjusted for this confounding variable in all subsequent statistical models. Species that differed significantly based on isolation method can be found in Supplementary Table 5.
Sequencing
Total genomic DNA was extracted from a fecal PBS aliquot using the MagAttract PowerMicrobiome DNA/RNA kit (Qiagen). Genomic DNA was then prepared for Whole Genome Sequencing analysis using the KAPA HyperPlus Library Prep™ kit (Roche). Sequencing analysis was conducted on the Illumina platform using paired-end 150 bp reads. For samples obtained from DNA/RNA Shield Fecal Collection Tubes, the DNA was extracted and purified using the Zymobiomics DNA Miniprep Kit. Extracted DNA was stored at −20 °C prior to use in whole Genome Sequencing analysis. Sequencing data was processed to remove low quality reads and adapter contamination using Trim-galore89.
Microbial classification
To identify the relative abundances of species in the infant fecal samples, a custom Kraken2/Bracken90,91 index was built specific to the gut microbiome. First, 1085 gut and oral genera were identified through Unified Human Gastrointestinal Genome (UHGG)92 and the MGnify human-oral database (http://ftp.ebi.ac.uk/pub/databases/metagenomics/mgnify_genomes/human-oral/v1.0/).
Subsequently, we analyzed our in-house large cohort of adult and infant gut microbiomes to identify additional genera not present in the UHGG and MGnify microbiome databases. This analysis identified 14 additional genera. Using GTDB taxonomy r207, we identified 17,752 unique species belonging to these genera. GTDB also provided assembly statistics for each genome corresponding to these unique species.
We downloaded 132,128 bacterial and archaeal genomes from NCBI, corresponding to our predefined unique oral and gut microbiome species, and selected those that met our quality criteria of low contamination and high genome completeness, corresponding to our predefined unique oral and gut microbiome species. These genomes underwent Mash-based clustering to dereplicate the assemblies93. We identified an optimal Mash threshold internally to retain strain-level information while removing nearly identical duplicate genomes. (Metagenomics-Index Correction software: https://github.com/rrwick/Metagenomics-Index-Correction).
Relative abundances were computed using the Bracken method90. Mock communities were simulated to validate this classification method, demonstrating increased accuracy at the subspecies level compared to the standard GTDB classification. The classification noise for this custom classifier was measured to be 0.5%. Within each sample, only taxa with a relative abundance of ≥0.5% were considered detected, which reduces the false positive rate for species detection but also reduces true positive detection for low abundance species.
Statistics and reproducibility
When using parametric approaches to data analysis, the centered log-ratio (CLR) transformation was applied to species relative abundance after imputation of zero values using multiplicative replacement. All statistical analyses were performed in Python using either the SciPy v1.10.1 ( or StatsModels ( or StatsModels v0.14.2 ( package. All p-values were adjusted for multiple comparisons using the Benjamin-Hochberg procedure (FDR).
A chi-squared test was performed to analyze associations between two categorical variables (Fig. 1D). A generalized linear model (GLM) was used to investigate associations between two taxonomic variables while controlling for covariates, such as those described in Fig. 1G.
Analysis of covariance (ANCOVA) was employed for Fig. 1E, as well as for the human milk oligosaccharides (HMO) and carbohydrate-active enzymes (CAZY) analyses. This allowed us to control for covariates, such as the method of sample processing (PBS versus DNA collection and isolation). Additionally, ANCOVA was used to examine taxonomic differences between samples processed with PBS versus DNA/RNA isolation, identifying 18 features that were statistically different. However, none of these were among the key microbial taxa highlighted in this paper or present in particularly large numbers (Supplementary Table 5).
For non-parametric approaches to data analysis, we used feature relative abundances. The Mann-Whitney U test was applied to assess associations between two groups (Figs. 1F, 4A, C). Spearman correlation analysis was used to examine associations between two continuous variables (Fig. 4B, D).
For Fig. 6C, a GLM model was used to investigate the association between pediatrician-validated adverse outcomes by two years of age and two independent variables: initial DMM cluster assignment and antibiotic exposure within the first two years. Antibiotic exposure was included as a covariate to control for its potential confounding effect. Relative risk (RR) was calculated from the GLM coefficients by first extracting the coefficients and determining the baseline probability using the model intercept. Odds ratios (OR) were obtained by exponentiating the coefficients and subsequently converted to RR using the formula:
$${{\rm{RR}}}=({{\rm{OR}}}\times (1-{{\rm{Baseline\; Probability}}}))/(1-({{\rm{Baseline\; Probability}}}\times {{\rm{OR}}}))$$
The GLM analysis revealed an association between DMM cluster membership and pediatrician-validated adverse outcomes. Because B. longum and B. breve were among the primary taxa driving the composition of these clusters, we sought to disentangle whether the observed associations were attributable to these specific taxa or to broader emergent properties of the microbial communities captured by the clusters. To address this, we employed Generalized Estimating Equations (GEEs), which more appropriately account for the correlated structure of microbiome data and allow for testing of individual taxa-level associations. Using this approach, we specifically assessed whether (1) infant-associated Bifidobacterium, and (2) B. longum and B. breve, were independently associated with adverse outcomes. For Fig. 6D, we used a feature selection pipeline for logistic regression analysis to identify key gene clusters associated with adverse outcomes while controlling for antibiotic use in early childhood as well as DNA collection and isolation. The pipeline follows three stages: (1) L1 Regularization (Lasso) with 10 repeated cross-validation to identify consistently important features; (2) Recursive Feature Elimination (RFE) on the top 50 consistently important features to further refine the selected features; and (3) Final Model Building using the top features identified in earlier stages.
Functional analysis
Functional capacity of the infant gut was identified using gene annotations provided by the UHGG database92. We mapped paired-end reads to a catalog of 11,019,595 microbial genes, quantifying gene level relative abundance using a combination of exact k-mer matching and an Expectation Maximization (EM) algorithm to handle multi-mapped reads.
The EM algorithm probabilistically assigns each read to its most likely gene of origin based on initial mapping probabilities. This involves iteratively updating these probabilities to maximize the likelihood of the observed read distribution, taking into account the current estimates of gene abundance levels. Through this process, the EM algorithm refines the assignment of multi-mapped reads, improving the accuracy of gene-level abundance estimates. Within each sample, only genes with an expected read count of ≥3 were considered detected.
After assigning reads to genes, we aggregated the gene-level abundances by functional annotation of choice such as human milk oligosaccharide (HMO) genes, carbohydrate-active enzymes (CAZymes), antimicrobial resistance genes, and virulence factors.
HMO Identification and quantification
We focused on six well-defined human milk oligosaccharide (HMO) gene clusters identified in the genome of Bifidobacterium longum subs. infantis (H1, H2, H3, H4, H5, and Urease). To account for genomics variation in HMO-related genes among different bacteria beyond Bifidobacterium, we focused on the KEGG Orthology (KO) group and protein product name for each of the 59 genes found in these six gene clusters, as provided by our functional classifier.
Due to HMO enzymatic promiscuity and the fact that some individual genes within the HMO clusters can share the same KO ID, we also included gene number thresholds for each gene cluster. This allowed us to determine whether a set of HMO gene clusters and their genes were present. If a gene cluster met our predefined thresholds (H1: 8/20 genes present; H2: 2/4 genes present; H3:¼ gene present; H4: 7/12 genes present; Urease: 8/12 genes present), we considered the gene cluster to be present. We then counted the individual KO gene counts and divided by the total number of identified classified genes in the sample to obtain relative abundance.
The gene thresholds were determined by running a subset of baby biome samples (n = 80) through a DIAMOND HMO pipeline94. The HMO database was built using proteins from the six well-defined HMO gene clusters identified in the genome of Bifidobacterium longum subs. infantis. We examined histograms representing the distribution of gene presence within each gene cluster and used internal domain knowledge to confidently select our predefined thresholds.
To identify the main taxonomic contributor of HMO genes for a given sample, we traced each KO and/or protein product name back to its gene of origin. This enabled us to link the gene to its corresponding taxonomic genome. We then determined which taxa contributed the most HMO genes and identified these are our top taxonomic contributors.
CAZy enzyme identification
To determine the relative abundance of sialidase and fucosidase enzymes in our metagenomes, we obtained the CAZy data95 specifically using the file available at ( This file enables us to link the Enzyme Commission number (EC number) to our functional database. Subsequently, we calculated the relative abundance of all identified CAZy genes and selectively filtered them by the enzyme column to isolate relevant enzymes (sialidase and fucosidase).
Antibiotic resistance gene identification
All microbial genes in our functional classifier were screened for antibiotic resistance using the DIAMOND program to perform a BLASTX-type search against the NCBI antimicrobial resistance database. The DIAMOND outputs were parsed and filtered to collect top hits for each gene. Considering positive hits with (i) E-values ≤ 1e-50 and (ii) bitscores >500. We used this information to flag genes that conferred antimicrobial resistance in our functional database.
Virulence Factor Gene Identification
All microbial genes in our functional classifier were screened for virulence factors using the DIAMOND program to perform a BLASTX search against the virulence factor full database (VFDB)96. The DIAMOND outputs were parsed and filtered to collect the top hits for each gene, considering positive hits with (i) amino acid percent identity >95% and (ii) bitscores >500. We used this information to flag genes that conferred antimicrobial resistance in our functional database.
Metabolomics
Targeted metabolomics was performed at Precion Inc. ( using a microbiome panel based on isotopically labeled internal standards. 157 samples were submitted for metabolomics analysis, 109 of which were from breastfed infants. Samples were provided as PBS aliquots and then dried to determine a mass for normalization. Samples were then homogenized and proteins precipitated. Extracted metabolites were evaluated using a Sciex Exion LC/Sciex 5500+ Triple Quadrupole Mass Spectrometer LC–MS/MS system and four different methods. Aliphatic organic acids, aromatic organic acids, and other negatively charged analytes were evaluated in negative mode using C18 reversed phase chromatography. Amino acids, amines, and other positively charged analytes were evaluated in positive mode using C18 reversed phase chromatography. Short chain fatty acids and lactic acid were derivatized with a substituted hydrazine and analyzed in negative mode using C18 reversed phase chromatography. Phenolic and indole metabolites were derivatized with a substituted sulfonyl chloride and analyzed in polarity switching mode using C18 reversed phase chromatography. Using these four methods, a panel was evaluated that includes: 2-methylbutyrate, 3-hydroxybenzoate, 3-hydroxyhippurate, 3-hydroxyphenylpropionate, 3-methylindole, 4-ethylphenol, 4-ethylphenylsulfate, 4-hydroxyphenylacetate, 4-hydroxyphenylacrylate, 4-hydroxyphenyllactate, 4-hydroxyphenylpropionate, acetate, agmatine, arginine, benzoate, betaine, butyrate, cadaverine, carnitine, chenodeoxycholate, cholate, choline, cinnamoylglycine, citulline, deoxycholate, enterodiol, enterolactone, glycochenodeoxycholate, glycocholate, hexanoate, Hippurate, imidazole propionate, indole, indole-3-acetamide, indole-3-lactate, indole-3-propionate, indoleacetate, indoleacetylglycine, indoleacrylate, indoleacrylglycine, indoxyl sulfate, inosine, isobutyrate, isoleucine, isovalerate, kynurenate, kynurenine, lactate, leucine, lithocholate, lysine, N-acetylserotonin, ornithine, p-cresol, p-cresol glucuronide, p-cresol sulfate, phenol, phenol glucuronide, phenol sulfate, phenylacetate, phenylacetylglutamine, phenylacetylglycine, phenylalanine, phenyllactate, phenylpropionate, phenylpropionylglycine, phenylpyruvate, propionate, putrescine, serotonin, thiamine, trimethylamine, tryptamine, tryptophan, tyramine, tyrosine, ursodeoxycholate, valerate, valine.
Microbe-metabolite network construction
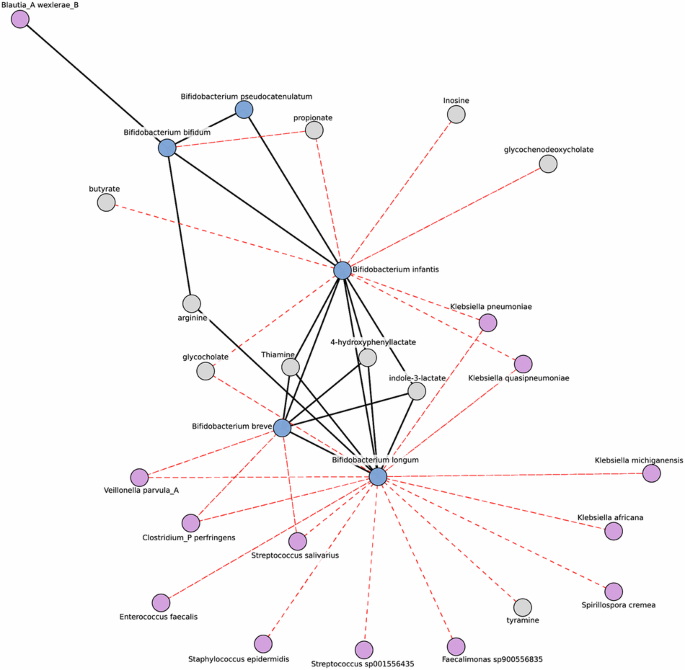
Microbe-metabolite co-occurrence networks were constructed using taxonomic and metabolomics features from 109 breastfed infants. Initially, taxonomic and metabolomics data were treated as independent data frames. The counts were transformed to relative abundances. Following this, center log ratio (CLR) transformation97 was applied to the data after imputing zero values using multiplicative replacement98 to address issues of compositionality. Finally, the two data frames were merged for subsequent analysis. Correlations and associated p-values were obtained using the Pearson correlation method and p-values were adjusted for multiple comparisons using the Benjamin-Hochberg procedure (FDR)99.
A network consists of nodes and edges, where nodes represent individual features and edges represent the connections between those features. This network contained 211 features and 724 connections. The undirected graphical network was generated by estimating a sparse inverse covariance (precision) matrix with L1 regularization using GraphicalLassoCV (https://scikit-learn.org/stable/modules/generated/sklearn.covariance.GraphicalLassoCV.html).
By examining the precision matrix, we can discern genuine correlations from spurious ones, as it highlights conditional dependencies among the variables. GraphicalLassoCV uses cross-validation to select the optimal regularization parameter, ensuring the model is neither overfitted nor underfitted and generalizes well to unseen data. The L1 regularization introduces sparsity, forcing many estimated connections to be zero, resulting in a simpler and more interpretable network. This is especially important for high-dimensional data, as it reduces the risk of overfitting and enhances network interpretability. NetworkX was used to visualize the graphs from the resulting output (https://networkx.org/documentation/stable/index.html).
Protein gene clustering using Markov Clustering Algorithm
Knowledge-based approaches (i.e., KEGG, COG, Gene Ontology) leave a large portion of predicted genes uncharacterized and naïve approaches provide an overwhelming amount of data that is difficult to parse, so we developed an ortholog clustering pipeline that allows gene families to be grouped together to increase their significance in a data set. This allows uncharacterized genes to be analyzed while still providing a manageable framework.
We converted 11,019,595 microbial genes from our functional database into an amino acid FASTA (FNA) file. From this file, we generated another FNA file by removing gene duplicates. This deduplicated FNA file was then used to build a DIAMOND database. We employed an all-versus-all DIAMOND approach, where the full FNA file was queried against the DIAMOND database, similar to the method previously described by Harlow et al.100. We focused on three DIAMOND output columns: qseqid, sseqid, and evalue. The data were parsed and filtered further, then passed through a Markov Clustering Algorithm (MCL) using the -abc-neg-log10 transformation101.
Ther MCL works with the input similarity file by transforming it into a matrix representing a graph. It then uses a series of matrix operations (expansion and inflation) to discover clusters within each graph. Each protein or gene sequence is treated as a node in a network, with edges representing sequence similarities based on the -log10(E-value). This transformation results in a symmetric matrix where the edge weights signify similarity: higher weights indicate more similar proteins, while lower weights indicate less similar ones.
Clusters are characterized by many heavily weighted edges between members, forming tight-knit groups with shorter paths (edge lengths) within a cluster. Conversely, proteins with no similar counterparts form single-protein and multi-protein clusters with long paths to other nodes in the network. The MCL algorithm helps us construct these networks and probabilistically identify protein gene clusters, resulting in a final matrix that can be interpreted as protein family clusters. The inflation value parameter of the MCL controls the granularity or “tightness” of these clusters.
This protein clustering approach has been shown to be effective at identifying protein families, including proteins with multi-domain structures and promiscuous domains (domains that are present in many families), with both internal and external validation102.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
link